Choosing Cancer Weapons With the Best Aim
In oncology, we do the best we can. We give drugs with 10-20% chances of working because we have nothing else. Drugs are approved, right or wrong, based on mere weeks of overall survival benefit. Clinical trials labeled as novel are merely the result of adding approved drug A to approved drug B and justifying it with a contrived methodology. We often use reasonably effective, but crude metrics, such as PD-L1 expression, mismatch repair (MMR) deficiency, microsatellite instability (MSI) status, and tumor mutation burden (TMB) levels to delineate if patients will respond to immunotherapy. Unfortunately, despite our best intentions, we are often wrong. Cancer cells don’t read a textbook. They don’t care about our predictions or our prognostic models. Accordingly, and I tell this to the people I teach all the time, what the cancer is doing matters far more than what you think it should do. Phenotype is ALWAYS more important than genotype. Nonetheless, it’s imperative we continue to strive to close the gap between expectation and actuality. Numerous companies, academicians, etc., have capitalized on our innate need for control as clinical practitioners and scientists. Artificial Intelligence (AI) and machine learning (ML) based drug response prediction models are being developed at exponential rates. Every week I’m approached by a new company claiming they can accurately identify patients who will respond to a particular drug, and those that will not. Today, we will bring together the short-term and long-term approach to clinical cancer care, AI/ML, multiomics, and cell-free DNA, covered previously in this series. In this eighth entry to “The Insider’s Guide to Translational Medicine”, we explore the future utility of drug response prediction models in the clinic through a case study of immunotherapy prognostic assays. Specifically, we will discuss Oncocyte and BostonGene, and provide the reader with an incredibly unique perspective UNAVAILABLE ANYWHERE ELSE.
Image credit: Adrienn Harto
You’re Back!
Despite your 2–3-month absence from the media, you now make your return as CEO of Comprehensive Precision (figure 1), the job you took after being part of the Biopharmatrend.com Investment Group (BPTCIG). While seated at your desk, proud of the command center you built for yourself (figure 2), you reflect on a request your board made a week ago to enter the treatment response prediction market.
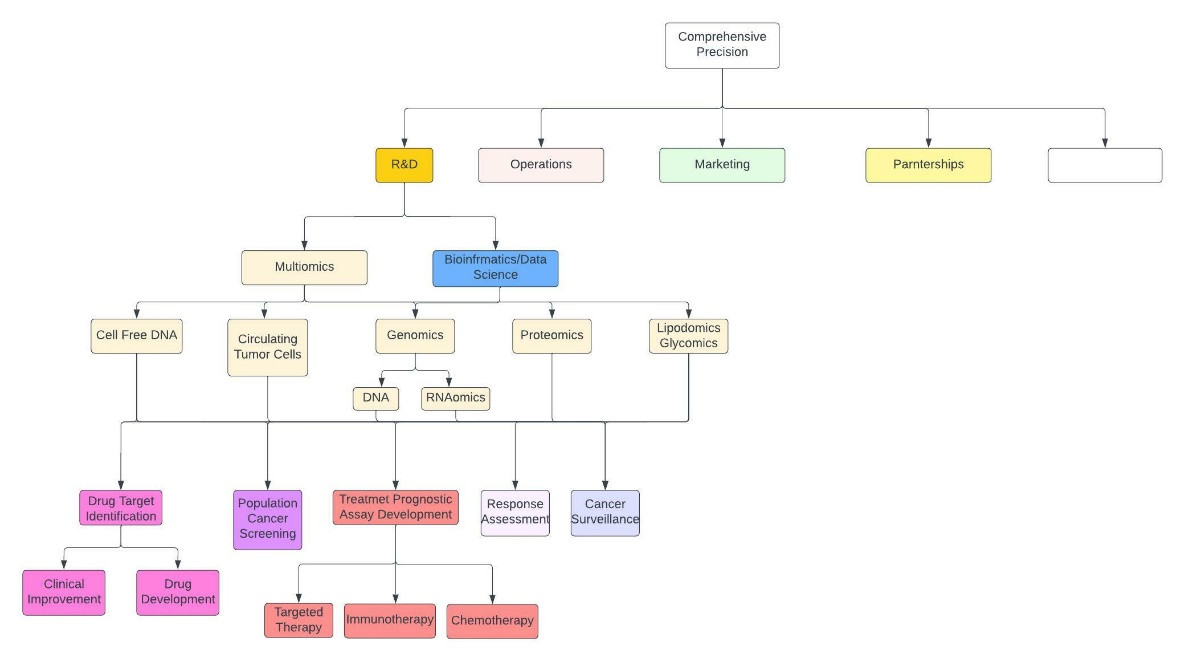
Figure 1: Abbreviated and crude description of Comprehensive Precision. The empty white box encompasses everything else.
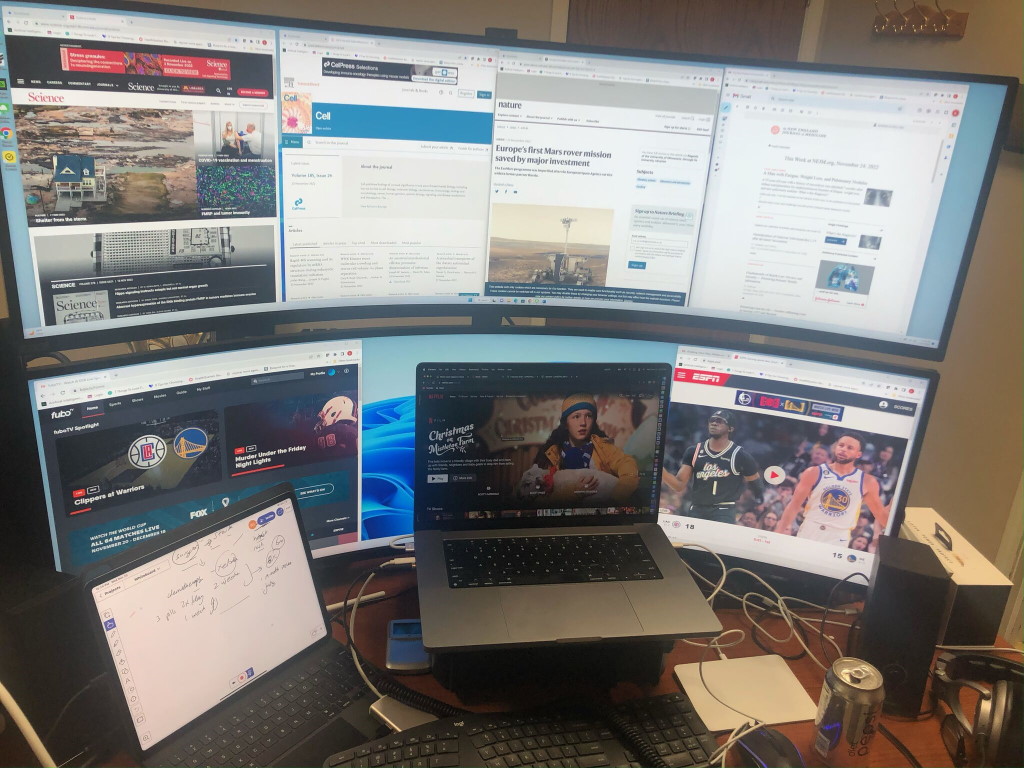
Figure 2: The command center in your office [2].
You’re aware your company’s primary core competency is multiomics, including proteomics, genomics (DNA and RNA), glycomics, etc. Yet, particularly in this financial climate, you’re extremely trepidatious about where you invest your company’s resources. While in Kuai, Hawaii (figure 3), as chronicled in prior entries to this series, you were forced by your board to propose a rudimentary strategic approach for entering the population health cancer screening market, despite your objections. Accordingly, it’s not surprising you board is now very keen on the use of multiomics in therapeutic response prediction, but you need to get there on you own. Indeed, you’re a bit annoyed that your board, largely comprised of people who know nothing about multiomics, medicine, or technology, talk as though they do. But you know it’s the world we live in; people don’t know what they don’t know, and they often talk as they do.

Figure 3: The view from your hotel, the “Cliffs at Princeville”, while you were in Kauai.
Therapeutic Response Prediction Overview
You are a voracious reader. You know there are innumerable companies developing therapeutic response assays. Indeed, the barrier to entry in the market is so low it doesn’t surprise you that, as a practicing hematologist/oncologist and CEO of Comprehensive Precision, new companies approach you weekly to use their drug response kits in the oncology clinic. What they don’t know is that you have been reading “The Insider’s Guide to Translational Medicine” series on biopharmatrend.com [1] and are now incredibly equipped to properly assess their offerings.
You note that immunotherapy, therapeutics designed to stimulate the immune system to eradicate cancer cells, has transformed hematology/oncology. Additionally, beginning with the CTLA-4 antibody, ipilimumab (yervoy), in melanoma, and continuing with numerous immune checkpoint therapeutics, the immunotherapy field is expanding so rapidly you can barely keep pace. Essentially every protein of the immune checkpoint signaling pathway (figure 4), including GITR, CD80, CD86, OX40, STING, TIM3, TIGIT, CTLA4, PD-1, PD-L1, ADO2, ADO1, CD73, CD94, Kir, A2aR, etc., is being targeted by pharmaceutical companies. Many of the corresponding drugs have fallen by the wayside. Others are in danger of doing so, including the TIGIT inhibitor, tirogolumab, which recently suffered setbacks in stage 4 non-small cell and extensive-stage small cell lung cancer. Fortunately, the PD-1 and PD-L1 inhibitors are firmly entrenched in cancer care and likely will be forever.
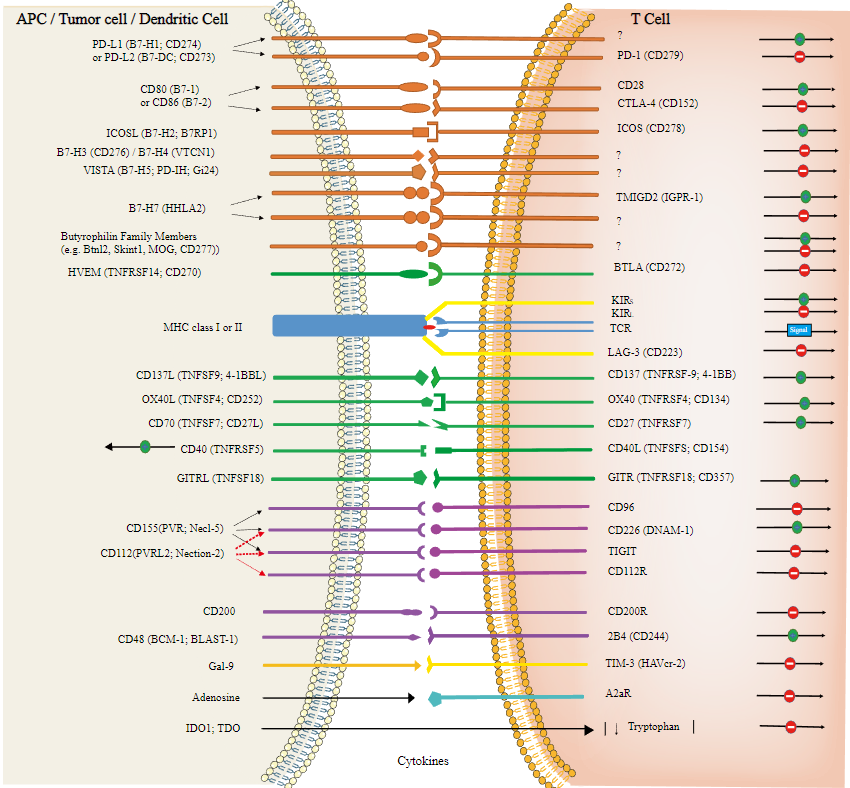
Figure 4: Beautiful illustration of the immune-checkpoint signaling pathway [3].
Presently, there are 5 FDA-approved PD-1 inhibitors, including pembrolizumab, cemiplimab, nivolumab, dostarlimab, and avelumab. There are 2 FDA-approved PD-L1 inhibitors, including atezolizumab and durvalumab. Numerous other PD-1 inhibitors are in development, including toripalimab, sugemalimab, sintilimab, tislelizumab, etc.
PD-1/PD-L1 inhibitors play a central role in nearly every cancer type, and prominently feature in lung, melanoma, bladder, renal, hepatocellular, head and neck, esophageal, gastric, etc., cancers. Interestingly, despite being used neoadjuvantly for triple-negative breast cancers, and in stage 4 triple-negative breast tumors that express PD-L1 in 10% or more of their cancer cells, PD-1/PD-L1 inhibitors have a minimal role in breast cancer treatment. Similarly, outside of patients with high levels of microsatellite instability (MSI-high) and/or tumor mutation burden (TMB-high), immunotherapy essentially has no current role in the treatment of metastatic prostate cancer.
You’re aware that current dogma is that patients whose tumors highly express PD-L1 have a higher chance of responding to PD-1/PD-L1 inhibitors. Moreover, patients with cancers that are mismatch repair (MMR) deficient, have high levels of microsatellite instability (MSI-high), and/or high tumor mutation burden (TMB-high) have a higher likelihood of responding to PD-1/PD-L1 inhibitors. Indeed, MSI-high status is such an excellent predictor of immunotherapy response, particularly in colorectal cancer, that dostarlimab and pembrolizumab are both FDA-approved for the tumor agnostic treatment of patients with MSI-high cancers.
Recently, a study involving 12 stage 2/3 rectal cancer patients who were MMR deficient (which generally correlates with being MSI-high) and treated with dostarlimab, revealed that all 12 patients had a complete remission. This enabled them to avoid standard treatment of chemoradiation, followed by surgery, followed by chemotherapy, or chemotherapy, followed by chemoradiation, followed by surgery. This study was remarkably over sensationalized as it was published in the New York Times (figure 5), in large part because it was conducted at Memorial Sloan Kettering (MSK), who has an excellent marketing team (4). To this end, all oncologists know that stage 4 colorectal cancer patients who are MSI-high or MMR deficient respond beautifully to immunotherapy. The NICHE-2 trial, reported at the ESMO Congress 2022 meeting (5), showed that 99% of 112 MMR-deficient, locally advanced, colon cancer patients, treated with pre-operative immunotherapy (1 dose of iplimumab and 2 doses of nivolumab) had a pathologic response, including 67% of which who had NO EVIDENCE OF RESIDUAL CANCER at the time of their definitive colon resection (Figure 6). Accordingly, the aforementioned MSK rectal cancer result was expected to a large extent, although the 12/12 complete remission rate is tremendous.

Figure 5: “A Cancer Trial’s Unexpected Result: Remission in Every Patient”, authored by Gina Kolata and published in the NYT, June 5, 2022 [4].
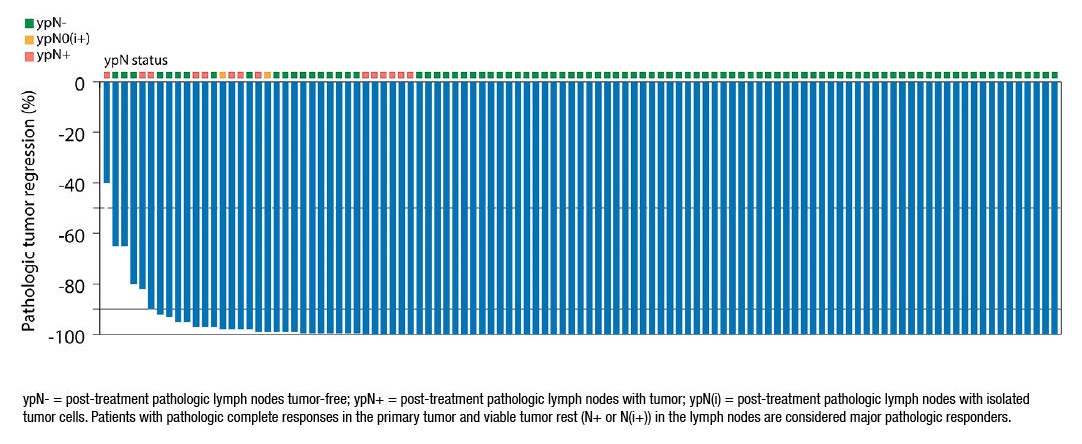
Figure 6: Results of the NICHE-2 trial showing the profound pathologic responses to pre-operative immunotherapy in MMR-deficient, locally advanced, colon cancer patients [5].
In your experience, oncologists, including yourself, use PD-1/PD-L1 inhibitors like water. The reason for this is predominantly because there are usually few viable treatment options, and they are relatively innocuous drugs. Although they always carry the risk of autoimmune related side effects, such as colitis, pneumonitis, nephritis, hepatitis, etc., these side effects are relatively infrequent.
Presently, our classic indicators of immunotherapy response, as above, are whether or not a cancer is PD-L1 high, MMR-deficient, MSI-high, or TMB-high. Stage 4 non-small cell lung cancer patients with tumors that are more than 50% PD-L1 positive are often treated with a PD-1 inhibitor alone, whereas tumors that don’t meet this criterion frequently require chemotherapy with immunotherapy. Stage 4 colorectal cancer patients that are MSI-high are generally treated with immunotherapy initially, as opposed to chemotherapy for cancers that are not MSI-high. In fact, essentially all patients that are MSI-high, irrespective of tumor type, will receive immunotherapy based on this.
Despite being better than nothing, there are ample times when patients one would expect to respond to immunotherapy based on high PD-L1, MSI, or TMB, or MMR deficiency, don’t. Moreover, patients one would not expect to respond to immunotherapy predicated on these factors occasionally do. Studies have shown that 1/6 patients with stage 4 NSCLC predicted to NOT respond to immunotherapy do [6]. Thus, you feel we’re in dire need of better immunotherapy prognostic markers in the clinic.
You recognize numerous companies are developing immunotherapy response prognostic assays. Many of them are being tested in the clinic as these companies are racing to be the first to have their tests approved by the FDA for commercial use. They recognize there is a TON of money to be made in this regard. This includes, but is not limited to, Oncocyte, BostonGene, and Oncohost (not formerly discussed here). Accordingly, although therapeutic response assays are being developed for almost every drug one can conceive of, in addition to immunotherapy, you elect to use them as your case study for the world of therapeutic response prediction.
Revealing Secrets
As CEO of Comprehensive Precision, you’ve been fascinated by how much money is being thrown at companies to develop rudimentary multiomics based tests. Oncohost just secured 35 million dollars in Series C funding after a series B round of 8 million. Oncocyte, which offers a DetermaRx stage 1A-2A lung cancer molecular stratification test, was once valued at 895 million dollars and is presently valued at 90.1 million. Boston Gene recently secured 150 million dollars in series B funding after a series A round of 50 million.
You reference the primer (Figure 7) you provided your board members in “Predicting a Patient’s Future with a Crystal Ball Comprised of Cell-Free DNA” [7]. In that primer you detailed how companies employ simple artificial intelligence/machine learning (AI/ML) techniques to develop population health screening assays. A data set comprised of cell-free DNA (cfDNA) corresponding to patients labeled as having cancer or healthy was used to create a training set and validation set to develop an AI/ML model that could predict if a patient had cancer based on their cfDNA. To this end, you recognize that developing therapeutic prognostic assays is based on the same principles.

Figure 7: Predicting a Patient’s Future with a Crystal Ball Comprised of Cell Free DNA [7].
Developing Your Own Therapeutic Response Assay
The data set
You are astutely aware that the primary difficulty of developing a therapeutic response assay resides in having the required data to do so. Specifically, you need access to a robust cancer patient population treated with the drug of interest, and their corresponding cfDNA, where patients in the data set are labeled as responders and non-responders. You then employ a supervised learning paradigm to build your treatment response prediction model (figure 8).
Building the model
The parent data set is separated into a training and validation set (figure 8). Thereafter, a treatment response prediction model is developed based on the training set and AI/ML techniques, such as logistic regression, principal component analysis (PCA), etc. (figure 9). Many model iterations are generated to optimize its ability to predict if a patient will respond to treatment or not, which is intrinsic to machine learning.
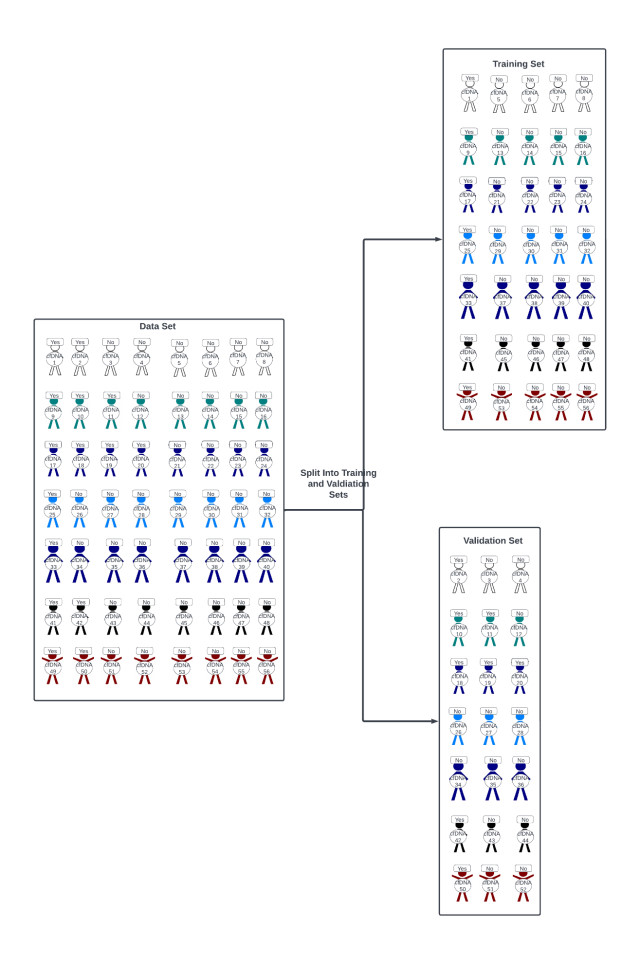
Figure 8: A data set corresponding to patients that responded to treatment and those that didn’t is split into training and validation sets.
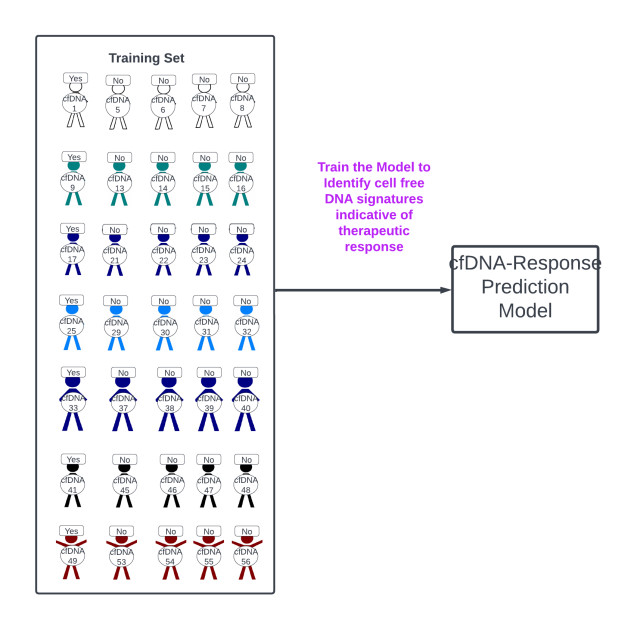
Figure 9: A treatment response prediction model is generated from the training set.
Validating the model
The response model developed using the training set is tested on the validation set to see if it accurately predicts patients who will respond to treatment (figure 10). Importantly, many people use training, validation, and test sets, but for your purposes you consider the validation and test sets one and the same. If testing reveals the model performs well on the validation set one is ready to take it out into the “real world”.
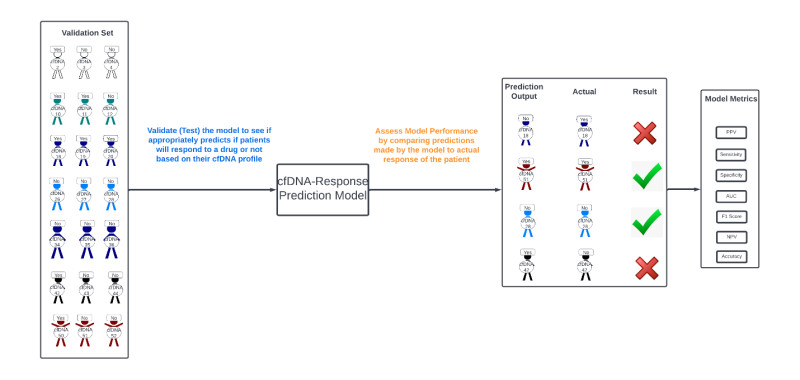
Figure 10: The generated treatment response prediction model is “tested” using a validation set.
Iterating through machine learning
It’s unlikely that one will generate an accurate treatment response prediction model on first pass. Indeed, such models are constantly being iterated to improve on their performance. The first model generated is almost never the final model used (figure 11). To this end, the more data one has the better model one will generally be able to develop. Consequently, you know there is a major war being waged across academic centers, pharmaceutical companies, precision medicine companies, etc. over patient data. Those with the data are the “haves” and those without are the “have-nots”. In fact, given that the development of models is relatively facile if one has the “necessary data”, you recognize having the “necessary data” is the primary determinant in who “wins” and “loses”, with the prize for winning being no less than potential billions of dollars.
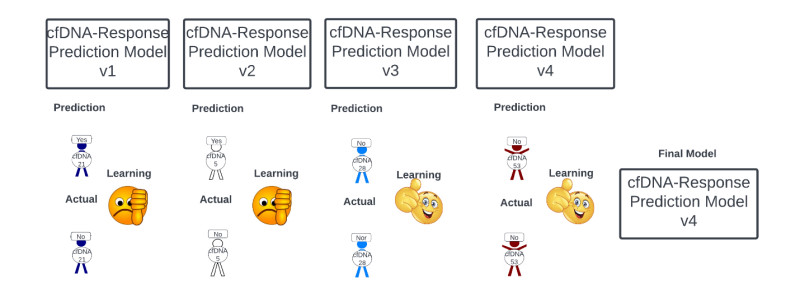
Figure 11: Macroscopic and rudimentary view of the machine learning process underlying the optimization of treatment response prediction models.
Launching the model
With final model in hand, you’re aware that aggressive marketing is employed to convince practicing physicians and patients to use the corresponding treatment response prediction assay. Indeed, the Holy Grail is that oncologists will ultimately administer patient’s treatment based on your model and its treatment response predictions (figure 12).
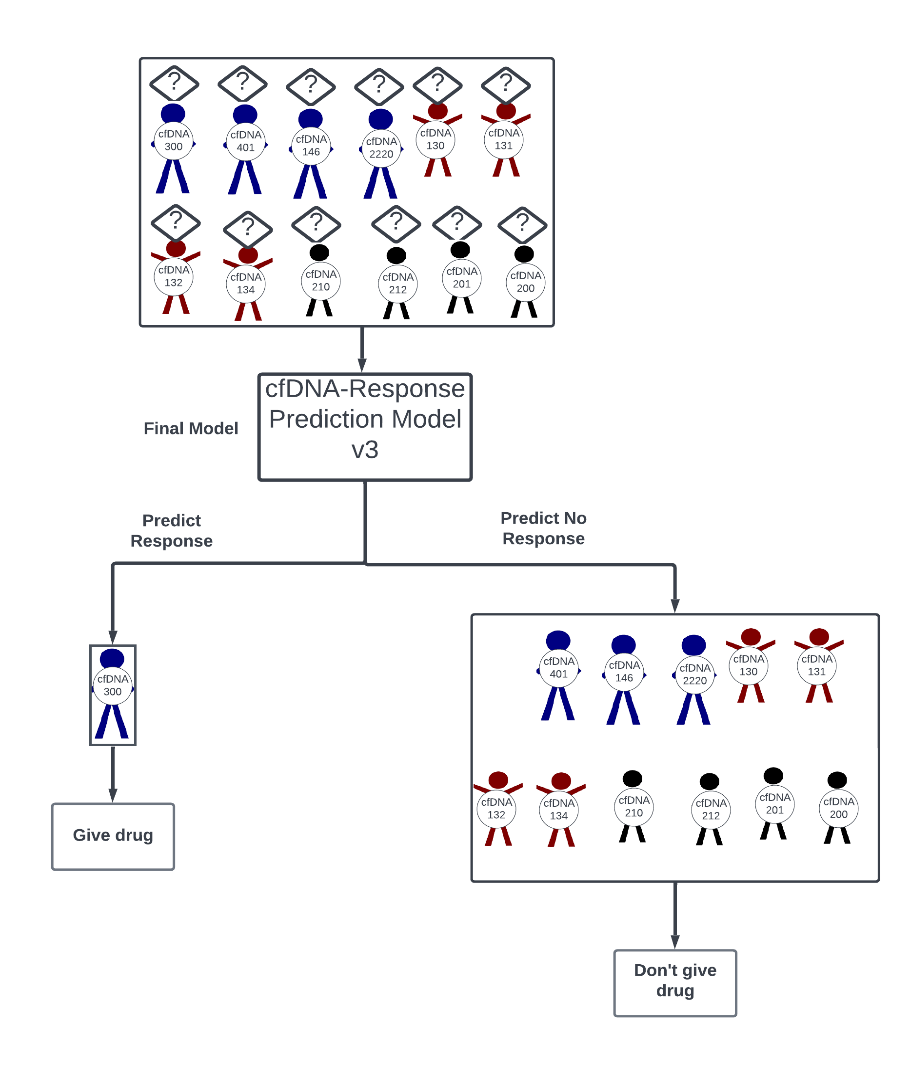
Figure 12: Use of the model in the “real-world”.
Real-world validation of the model
You’re irritated that many companies have been launching their models before they’ve been adequately validated in your opinion. Indeed, you believe that generating a therapeutic response model is relatively facile. Anyone can contend they can predict if someone will respond to a drug or not. The question is whether they’re right or not. Indeed, oncologists such as yourself are being inundated with companies claiming they can make you better at your job by dictating how best to treat your patients.
Nonetheless, you believe it’s imperative that therapeutic response assays be put through their paces to delineate their true utility. This can be done retrospectively but is best done in a prospective study. Ideally, one would conduct a study where patients were given a drug or not based on what the assay predicts and determine if patients treated accordingly have a better overall survival than those treated via status quo paradigms. However, you note there is profound risk in this. Specifically, you contend that what ABSOLUTELY CANNOT HAPPEN, is that a drug that would benefit a patient is withheld based on an incorrect prediction by the assay. In addition, you feel it would be a travesty if a patient lost time while being treated with a futile drug because a response assay incorrectly predicted they would respond to treatment. Accordingly, you believe treatment response predicting, although potentially very lucrative, is a very dangerous game. You better not be wrong!
Because of these considerations, initial real-world testing of therapeutic response assays doesn’t usually involve giving patients a given drug solely based on the response assay prediction. Generally, it entails comparing the response assay prediction to a patient’s response to treatment retrospectively. If the sensitivity (precision), specificity, positive predictive value (recall), etc., is high enough it may be deemed ready for prime time and potentially be approved for real-world use to guide whether a patient should receive a given drug, but this process isn’t ideal, as above.
Different response prediction assay methodologies
In our rudimentary example we developed a therapeutic response assay using cancer patient cfDNA. However, one could conceivably compare drug response to a patient’s protein, lipid, carbohydrate RNA, etc., profile. In addition, the fundamental nature of a company’s response assay will almost certainly differ from everyone else’s. Even if every company used cfDNA to predict drug response, one company may look at the upregulation or down-regulation of 27 genes, such as Oncocyte, or 100 genes. In addition, even if every company looked at the same number of genes, 27, they may look at different genes. And even if they used the same 27 genes, they may ascribe a different weight in their treatment response prediction algorithm to each of the genes than another company. Thus, it’s basically a certainty that no two companies will have the same response assay.
Determining the winners and the losers
Identifying the best drug response assays resides in confusion matrices (figure 13) introduced in the “Drowning in Data” entry in “The Insider’s Guide to Translational Medicine” [8]. Basically, one determines the number of true-positives, false-positives, true-negatives, and false-negatives associated with the assay of interest.
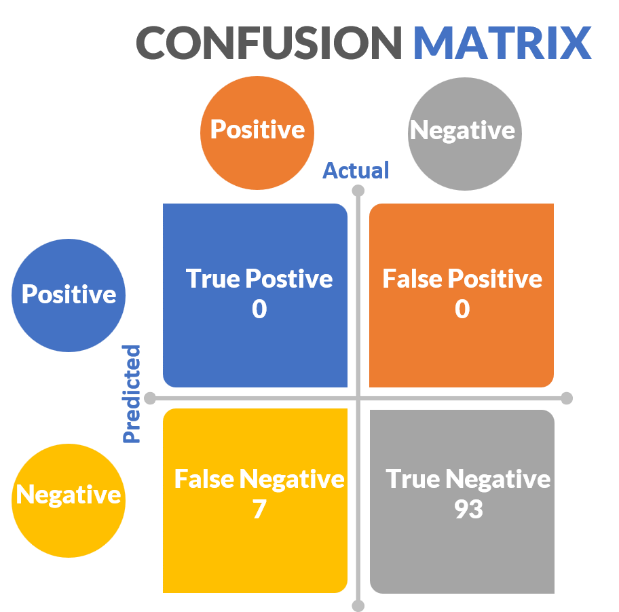
Figure 13: Hypothetical confusion matrix
It’s not complicated and distills down to basic metrics such as sensitivity (also known as recall), specificity, positive-predictive value (also known as precision), AUC, etc. (figure 14).
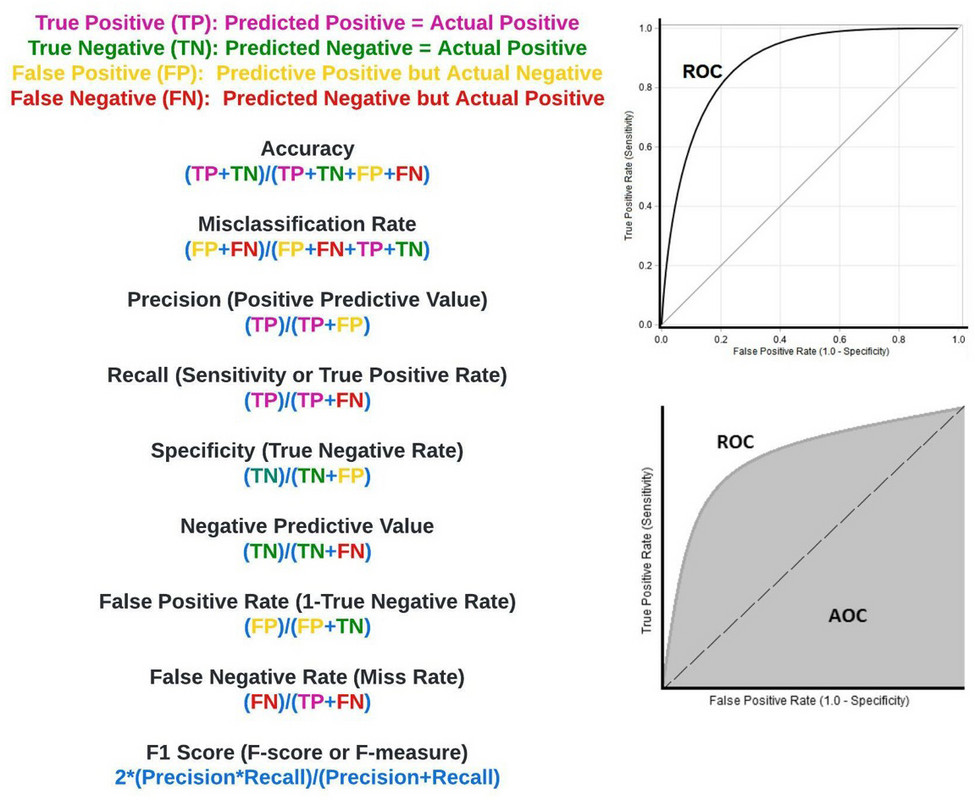
Figure 14: Confusion matrix associated metrics.
You now consider a hypothetical therapeutic response assay a company contends can correctly predict if patients that have stage 4 non-small cell lung cancer and are PD-L1 negative, TMB low, and MSI stable, will respond to keytruda. In this theoretical study exploring 100 such patients treated with keytruda the assay predicts 20 patients will respond to keytruda, and 80 will not. In reality, in the study, 9 patients respond to immunotherapy and 91 do not. Of the 9 patients who responded to immunotherapy the model correctly identified 4 of them. Of the 20 patients the model labeled as responders, only 5 responded to keytruda.
Building the associated confusion matrix (figure 15) true positives (TP), true negatives (TN), false positives (FP) and false negatives are as follows:
TP (4): Patients the assay correctly predicted would respond to keytruda
TN (75): Patients the assay correctly predicted would NOT respond to keytruda
FP (15): Patients the assay incorrectly predicted would respond to keytruda
FN (5): Patients the assay incorrectly predicted would NOT respond to Keytruda
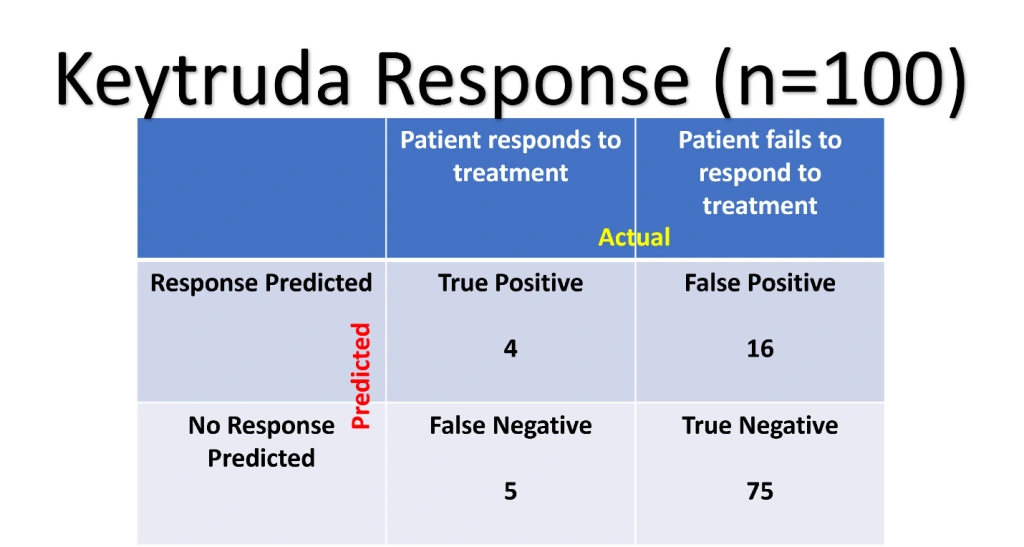

Figure 15: Confusion matrix corresponding to the Keytruda response prediction assay.
Now you calculate the confusion matrix metrics:
Accuracy: (TP+TN)/(TP+TN+FP+FN) = 79/100 = 79%
Sensitivity (Recall): TP/(TP+FN) = 4/9 = 44.44%
Specificity: TN/(TN+FP) = 75/91 = 82.4%
Positive Predictive Value (Precision): TP/(TP+FP) = 4/20 = 20%
Negative Predictive Value: TN/(TN+FN) = 75/(75+5) = 93.75%
You forego calculating AUC, F1-score, etc.
You forego calculating AUC, F1-score, etc.
Assessing the metrics, you note the overall accuracy was 79%, which en face isn’t terrible. Digging deeper though, you realize the sensitivity, the rate with which the model identified keytruda responders, was only 44.44%. The specificity, the rate at which the model correctly predicted patients will NOT respond to keytruda, was 82.4%. The positive predictive value, the rate at which patients the model says will respond to keytruda actually respond to keytruda, is 20%. The negative predictive value, the rate at which the model contended patients will NOT respond to keytruda who didn’t respond to keytruda, is 93.75%. You conclude these numbers aren’t great, but could be sold to the less educated investor.
You indulge this thought for a moment and ponder how you would sell this hypothetical Keytruda response assay to an unsuspecting investor or consumer. It’s clear that you would hide the sensitivity of 44.44% and the positive predictive value of 20%. You would emphasize the accuracy of 79%, specificity of 82.4% and negative predictive value of 93.75%. You smile for a moment as you’ve seen investors and companies duped with far worse antics in your career.
As illustrated in the “Drowning in Data” article you read previously, many companies seek investment by relying on their negative predictive value. They are able to do this in situations where there is profound data imbalance, even when their products are suboptimal. Specifically, in populations with a preponderance of true non-responders, the negative predictive value will be high by default. Accordingly, just as in this case, the true measure of the response assay is the sensitivity (recall) and positive-predictive value (precision). With this in mind you are reminded to ALWAYS ASK ABOUT PRECISION AND RECALL, which reveals the hypothetical model as atrocious.
Ultimately, with the aforementioned understanding, you begin assessing Oncocyte and Boston Gene.
Oncocyte
You note Oncocyte is farther along than its competitors in developing an immunotherapy response assay. Their DetermaIO test (figure 16) is comprised of a 27-gene set they claim can identify a tumor microenvironment (TME) signature predictive of immunotherapy response. In their seminal study, DetermaIO was studied in triple negative breast cancer patients who received chemotherapy and durvalumab immunotherapy in the neoadjuvant setting [9]. Specifically, the test was used to predict which patients would have a pathologic complete remission (pCR) at the time of surgery after receiving chemotherapy and durvalumab immunotherapy. DetermaIO predicted a pCR with a sensitivity of 65.2%, specificity of 68.8%, PPV of 60%, and NPV of 73%. CONVENTIONAL PD-L1 testing had a better sensitivity of 74%, but a lower specificity of 48%; PPV was 55% and NPV was 68%. Using DetermaIO and PD-L1 in concert had a sensitivity of 65%, specificity of 74%, PPV of 68%, and NPV of 72%.

Figure 16: Oncocyte’s DETERMAIO [10].
You note there were several limitations to this study. Initially, only 50 patients were assessed using DetermaIO. Second, durvalumab immunotherapy was given with chemotherapy prior to surgery as immunotherapy undoubtedly wasn’t solely responsible for pCRs. Regardless, at face value you feel the DetermaIO metrics are not that impressive. Basically, the test properly predicted patients who would have a pCR 65% of the time, but failed to do so 35% of the time. It properly predicted patients would not have a pCR 68.8% of the time, but was wrong 31.2% of the time. Simple PD-L1 testing was actually better at predicting which patients would have a pCR.
You note Oncocyte did test DetermaIO in 43 stage 4 non-small cell lung cancer (NSCLC) patients [11]. They reported an odds ratio of 5.76, but the 95% confidence interval was 1.3 to 25.51. Again, you feel it’s impossible to draw any conclusions from this study as it only included 43 patients, had a gigantic confidence interval, etc.
Amazingly, despite the aforementioned data and the sheer simplicity of the DetermaIO test, the company was valued at nearly a billion dollars in 2021; it’s presently valued at 90 million.
BostonGene
BostonGene developed the Tumor PortraitTM test, based on integrated genomic and transcriptomic analysis, to predict if a patient will respond to immunotherapy. They have numerous partnerships for this, including with WellDyne. Moreover, they have a strategic agreement with the National Comprehensive Cancer Network (NCCN) to integrate its clinical practice guidelines into the BostonGene Tumor Portrait Test reports.
BostonGene utilizes a decision tree machine learning algorithm called Kassandra that was trained on a broad collection of over 9,400 tissue and blood sorted cell RNA profiles incorporated into millions of artificial transcriptomes to reconstruct the tumor microenvironment (TME). BostonGene contends that identifying a cancer’s TME subtype delineates whether or not it will respond to immunotherapy. Accordingly, the BostonGene Tumor Portrait Test uses DNA whole exome (WES) and RNA transcriptome (RNAseq) sequencing to analyze the tumor and its TME to determine if a patient will respond to immunotherapy (figure 17).
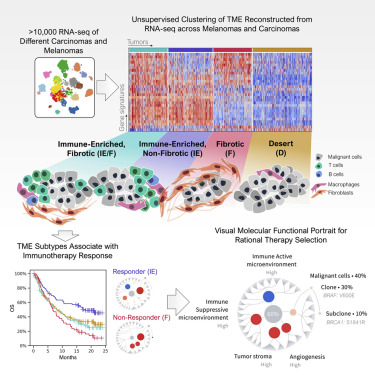
Figure 17: BostonGene’s Tumor Portrait Test [12] was based on over 10,000 RNA-seq from different carcinomas and melanomas (upper left). Unsupervised clustering based machine learning was performed, and generated 4 unique TME signatures (IE/F, IE, F, and D-middle). Patients with TME subtypes had varying overall survivals (lower left) and helped delineate immunotherapy responders and non-responders (TME IE subtype responded well, whereas TME F did not). A ”visual molecular functional portrait for rational therapy selection” is generated based on TME and other cancer related features, such as angiogenesis, tumor stroma, etc. (lower right).
The Tumor Microenvironment (TME) in brief
It is now well understood that the TME plays a central role in tumor response to therapeutics, particularly immunotherapy. To this end, our immune system is at constant war against aberrant cells that are malignant or can become malignant. It's constantly surveilling our cells to identify those that may give rise to various cancers, in a process called immunosurveillance, or trying to eradicate cells that are already malignant,
Unfortunately, the reason we need hematologists and oncologists is that in some cases cancer cells learn to evade the immune system cells. Current dogma is that this is often due to changes in the TME. Specifically, cells that inhibit immune system function can infiltrate the TME, including T-regulatory, myeloid-derived suppressor cells (MDSCs) and tumor associated macrophages (TAMs). In addition, alterations in immune checkpoint pathways central to immunosurveillance of aberrant cells (e.g PD-L1, TIGIT, VISTA, LAG3, CTLA4, TIM3, etc.), etc., can dramatically change the composition of the TME. Secretion of modified collagen into the TME can preclude cytotoxic T-cell lymphocytes, critical to immunosurveillance, from accessing aberrant cells. Release of inhibitory cytokines and molecules (e.g. adenosine) into the TME can also impede immune cell function. Consequently, numerous researchers, pharmaceutical companies, biotechnological companies, etc., are invested in elucidating the TME and developing associated treatments. BostonGene is foremost among them.
Back to BostonGene
BostonGene claims they can better predict if cancer patients will respond to immunotherapy better than the status quo by ascribing a patient’s cancer’s TME to one of four distinct TME subtypes based on genomic and transcriptomic data of the tumor and its TME (figure 17). They based their TME stratification on stromal, vascular, and cytokine expression patterns.
In their seminal 2021 Cancer Cell paper authored by Bagaev et. al. (12), they defined the aforementioned four TME subtypes based on unsupervised analysis of approximately 2,000 melanoma patients. They contended the four TME subtypes were conserved across at least 20 additional cancers in over 8,500 tumor samples, and TME subtypes correlated with patient overall survival and progression free survival when treated with immune checkpoint blockade (figure 17, lower left). Most importantly, they reported that they could predict cancer patient immunotherapy response to immune checkpoint blockade based on their TME subtype (figure 17, lower left).
This assertion was predicated on a study of samples from 58 patients with skin cutaneous melanoma treated with anti-CTLA-4 therapy (ipilimumab). Specifically, they found that 82% of patients with a non-fibrotic TME subtype they designated IE responded to treatment, whereas only 10% of patients with a different TME subtype they called F responded to treatment. They saw the same phenomenon in114 anti-CTLA-4 treated melanoma patients treated with anti-PD1 treatment as 75% of patients with the IE TME subtype responded to treatment, but only 10% of the F TME subtype did. 38% of 346 TME IE subtype bladder cancer patients treated with anti-PD1 or anti-PD-L1 therapy responded to treatment, whereas less than 10% of F subtype patients did. They observed the same patterns in 27 patients with lung cancer and 34 patients with gastric cancer.
In the Cancer Cell paper Bagaev et. al. contended that TME subtyping, central to BostonGene’s immunotherapy prognostic assay, performed better than traditional biomarkers of immunotherapy response, such as TMB, PD-L1, IFN gamma expression, etc., in predicting overall survival associated with immune checkpoint blockade in melanoma. Accordingly, they generated their Molecular Functional (MF) Portrait assay, which is their integrated transcriptomic and genomics based model for personalized tumor therapy selection. They state the assay incorporates malignant cell and TME characteristics, and is unique for each tumor type, despite there being 4 predominant TME subtypes observed across cancers.
The MF Portrait assay is presently being marketed, despite not being adequately validated in your opinion. In fact, there is no prospective trial showing that use of the assay to dictate cancer patient treatment is truly better than the status quo in any capacity. Nonetheless, you are personally aware that BostonGene is talking to insurance companies about incorporating their assay into their insurance paradigms.
You are extremely concerned about the ramifications of Boston Gene and insurance company conversations, including the fact that, as an oncologist, you don’t want an insurance company to utilize a non-validated assay to tell you how best to treat a patient. Indeed, you’re very concerned insurance companies will use such an assay to deny a patient a treatment that is FDA approved and NCCN supported. You cite the fact that, no matter what the BostonGene assay tells you, there is no way you’re not treating a stage 4 metastatic melanoma patient with immunotherapy, especially in the 60% of them that don’t have a BRAF V600E mutation. Accordingly, although it’s easy to sensationalize the BostonGene assay, you note that it has no current role in dictating treatment decisions in Oncology. To this end, you contend that BostonGene chose low-hanging fruit to test their assay on, but didn’t test their assay in the most challenging tumors to predict immunotherapy response. Your personally curious how it would behave prospectively in a large population of stage 4 non-small cell lung cancer patients who are PD-L1 negative and treated with chemoimmunotherapy. You would even be interested to see if the assay can properly predict which stage 4 non-small cell lung cancer patients that are more than 50% positive for PD-L1 will NOT respond to immunotherapy.
Collectively, although you note that BostonGene has done a tremendous amount of work to develop their assay and overall model, you’re not remotely ready to declare it a success. In fact, you note there is a large amount of money and politics behind BostonGene. They are intimately associated with MD Anderson as their CMO, Nathan Fowler, works there, and BostonGene recently had a series B round of $150 million. They partnered with NCCN to incorporate their guidelines in their assay reports. Accordingly, although there a lot of bells and whistles depicted on the BostonGene website and in the aforementioned Cancer Cell article, there isn’t a lot of substance to go by as yet in your opinion.
A quick clinicaltrials.gov search revealed that BostonGene is running its own patient registry to get access to patient samples to further develop their platform. They are looking for 10,000 patients and have connections with numerous large name institutions, as aforementioned.
To enter or not to enter? That is the question.
After your cursory review of the treatment response prediction market through assessment of immunotherapy prognostic assays you would prefer not to enter the market. You feel your company, Comprehensive Precision, would be far behind its competitors, and simply doesn’t have access to the patient data required to expeditiously build a treatment response prediction algorithm. To this end, you are painfully aware that the money in precision medicine isn’t in the tests themselves, it’s in the data acquired from performing the tests. You note that Illumina and AstraZeneca announced a NEW collaboration to identify new drug targets through genomic screening of patients. GSK announced they will pay Tempus 70 million dollars for access to their huge patient data repository containing genomics information, pathology images, etc. However, you are painfully aware that it takes data to get data. Indeed, without a model there is no test to obtain data with, and without the data there is no model.
It’s for this reason that every precision medicine company is racing to partner with institutions to get access to patient data. Oncocyte, BostonGene, and Oncohost (not formally discussed here) have established numerous relationships in this regard, and will continue to try to expand their networks.
As a new company you do not feel you can compete for patient data with these companies and elect to forego entering the treatment response prediction precision medicine market. In fact, big name academic institutions are now selling their data to the highest bidder. MSK, Dana Farber, Vanderbilt, Mayo, etc., have all sold the rights to patient data to companies.
You find this oft putting as these institutions are making money coming and going. Taxpayers provide funding for grants and their samples to such institutions only to unknowingly find that these institutions then develop assays and drugs with this money that are ultimately sold back to taxpayers. In addition, they sell taxpayer data and access to their biological samples to companies to line their pockets further, without taxpayers formally consenting to this. Vanderbilt recently sold access to their biobank to Illumina, and you’re fairly certain patients had no idea this was in the realm of possibility when they consented to have their samples donated to the biobank.
Accordingly, you’ve concluded precision medicine, like so much in life, is a forum where the rich get richer and the poor get poorer. In such a world, it’s very difficult for a new entrant to penetrate the treatment response prediction market, no matter how talented they are. The playing field simply is not level and numerous big players are incentivized to ensure it stays that way.
Wars in academia, the pharmaceutical industry, the precision medicine space, etc., are ALREADY being waged in the data battlefield, and this will only intensify. As such, you conclude that Comprehensive Precision will forego the treatment response prediction market as it doesn’t have adequate weaponry to enter the data battlefield.
Concluding Remarks
After performing your rudimentary analysis of the aforementioned companies your head hurts a bit. You take a moment to reflect and delineate the true utility of the aforementioned tests in the clinic.
You note that immunotherapy prognostic assays are likely to be expensive. Accordingly, for them to have true utility they should be more than just glitz and glamour, or a way to steal money from unsuspecting investors. They need to be a marked improvement over the status quo, which includes using PD-L1, MMR, MSI, and TMB to predict if a patient will respond to immunotherapy. In your opinion none of the aforementioned immunotherapy prognostic assays have definitively proven they are substantially better than the status quo in a RETROSPECTIVE manner, let alone a PROSPECTIVE ONE. Indeed, you would require clinical trials involving thousands of patients before you would be remotely convinced such assays will be useful to you clinically. You equate this to population health precision medicine companies studying their assays in tens of thousands of patients, including Freenome (35,000), Grail (150,000), etc.
As an oncologist you’re aware that the idea of using transcriptomics, proteomics, etc., isn't too different than tests that have been used in the clinic for years. For example, the Oncotype DX assay, which surveys 21 genes in a patient’s tumor, has been used extensively to determine if patients with early-stage breast cancer merit chemotherapy or not. Mammaprint, used for the same purpose, surveys 70 genes in the patient’s tumor. Accordingly, there is precedent for using -omic based assays to predict response to treatment. However, before immunotherapy response assays are used in the clinic they need to be far better validated than has been reported to date.
One in 6 stage 4 non-small cell lung cancer patients, predicted to be immunotherapy non-responders because they are not PD-L1 positive, TMB-high, or MSI-high, respond to immunotherapy. This means one would expect 17% of such patients to respond to immunotherapy and 83% not to. Therefore, in the absence of any immunotherapy prognostic assays one would be correct 83% of the time if they simply predicted the patient wouldn’t respond to immunotherapy. For you to be a believer of the aforementioned assays, they would need to be able to predict to a “reasonable degree of certainty” who the 17% of patients that will respond to immunotherapy are.
Determining what this “reasonable degree of certainty” is becomes far more challenging. The underlying premise is that you cannot afford to deny a patient a life-saving drug based on an incorrect prediction they will not respond to it. Conversely, treating a patient with an ineffective drug based on an incorrect prediction can be catastrophic as a patient may lose time and “ground” on the wrong treatment. Therefore, how good is good enough?
Are we willing to make treatment decisions based on assays that have 90% specificity for predicting what PD-L1 negative, stage 4 non-small cell lung cancer patients will respond to immunotherapy? What about the other 10% of such patients that would have responded to treatment that were denied the drug based on an assay with 90% specificity? What if that patient was your mother, father, brother, sister, or child? Are you content with them being collateral damage of these assays so insurance companies can save money by requiring oncologists to make immunotherapy decisions based on an unsupervised algorithm with minimal explainability and intuition?
You equate treatment response prediction to predicting winners and losers in sports. You note that in sports there are favorites and there are underdogs. Sometimes there are “heavy favorites” one can hardly surmise will lose a given match. Yet, the game still gets played. The sport still goes on.
We don’t hand out trophies at the beginning of a sports season based on how great a team looks on paper. The Saudi Arabias of the world do sometimes beat the Argentinas in World Cup soccer, no matter how inconceivable it may seem. Buster Douglas does occasionally knock out Mike Tyson. The US hockey team sometimes defeats the Soviet Union for gold in the Olympics. The New York Giants of the world occasionally beat the undefeated New England Patriots in the SuperBowl.
It’s why we play we play the game in the first place, no matter how obvious the outcome may appear…
Accordingly, in so many cancer patients with end-stage malignancies where treatment options are extremely limited, are we really not going to play the game? Are we going to admit defeat based on some unexplainable algorithm that hasn’t been tested prospectively solely because it predicts with 90% specificity a patient will not respond to a treatment? What if we already know that, based on pure statistics, 83% of such patients will not respond to the drug?
At this time, as a full-time hematologist/oncologist and CEO of Comprehensive Precision you are not prepared to declare any of the aforementioned assays a definitive solution to what ails us. They haven’t been validated enough to change your current management of cancer patients, and you need to see a LOT more before I accept them as gospel or anything remotely close to it.
You are painfully aware of the money investors have poured into the multiomics field. You’re acutely cognizant of all the companies rushing into the therapeutic response prediction space due to the minimal barriers to entry, relative simplicity, etc. Indeed, your contention remains that the secret sauce ISN’T in the algorithm, it’s in having access to the data.
You fully expect that the future of oncology will involve treatment response prediction assays, but you are leery of how this will affect patient’s lives. As you shut down your computer and walk towards the door of your office, you have one final thought as you turn off the lights…
Choosing cancer weapons with the best aim is everything in oncology, but what the cancer does matters far more than what you predicted it would do…
References:
1. https://www.biopharmatrend.com/topic/the-insiders-guide-to-translational-medicine/posts/
2. https://www.linkedin.com/posts/basem-goueli-md-phd-mba-16347810b_today-i-am-thankful-for-bosses-who-were-so-activity-7001529240529559552-gb5f?utm_source=share&utm_medium=member_desktop
3. https://www.creative-diagnostics.com/immune-checkpoint-signaling-pathway.htm
4. Gina Kolata, NYT, June 5, 2022
6. Khagi et al. (2017) Next generation predictive biomarkers for immune checkpoint inhibition. Cancer Metastasis Rev 36:179.
8. https://www.biopharmatrend.com/post/523-remember-to-be-precise/
9. Iwase et. al. A Novel Immunomodulatory 27-Gene-Signature to Predict Response to Neoadjuvant Immunochemotherapy for Primary Triple-Negative Breast Cancer. Cancers. 2021. 13(19). 4839.
10. oncocyte.com
11. Nielsen et al. A novel immuno-oncology algorithm measuring tumor microenvironment to predict response to immunotherapies. Heliyon. March 2021; 7(3): e06438.
12. Bageav et. al. Conserved pan-cancer microenvironment subtypes predict response to immunotherapy. Cancer Cell. June 14, 2021. 39(6): 845-865.