Pharma’s Data Dilemma: How Machine-Learning Tools Hold the Key to Unlocking Novel Insights
Covid-19 is still having a devastating effect on many countries and economies worldwide and has brought to light the impact of diseases that affect some people in worse ways than others. With SARS-CoV-2, ethnicity, gender, age, and underlying health conditions all play a part in disease severity and mortality, but there is mounting evidence to suggest that the status of the immune system early after infection can be predictive of those who go on to have the worst symptoms. The scale and threat of the pandemic has led to the need for rapid analysis of the disease – because the better we understand it, the more easily we can help healthcare professionals to make critical personalised treatment decisions. Advanced technologies such as artificial intelligence and machine learning are playing a large role in this process, and with other diseases, too, accelerating scientific research to enable major breakthroughs.
Digital first for pharma
The life science industry has been talking about artificial intelligence and machine learning for a long time and the benefits of increasing throughput, reducing human error, boosting productivity, and saving time and money have helped to cement automation within pharma and biopharma R&D. However, the extensive amount of high-quality data available within drug development pipelines poses a significant challenge for scientists trying to derive insights.
The range of different techniques being used generates data in many different formats and sizes, creating difficulties when structuring, sharing, and analysing the research. Data analysis is often still highly manual and requires specialist programming and data science skills, and it has been estimated that two thirds of researchers’ time is spent on processing data – valuable time that could be used for higher value scientific analysis and the complex tasks that researchers do best. For R&D labs to gain maximum value from data in a realistic timeframe, data processing and analysis technology must keep pace with automation innovation.
The value of flow cytometry in pharma research
Flow cytometry is a diverse and crucial technique in pharmaceutical research, used to investigate disease aetiology and alterations in immune responses, as well as for quantitative studies. There are several different steps during the flow cytometry data life cycle which include: data acquisition, processing, population selection, results integration, data analytics and insight generation. Unfortunately, its high throughput, multiparameter functionality is hampered by this immense output of highly complex data, especially with modern equipment. It requires significant expertise to interpret the data correctly, and there is a lack of standardisation in assay and instrument set-up. The technique has the potential to be used in every stage of drug discovery and development, so making it more efficient could have major, positive consequences for big pharma.
Flow cytometry data analysis is built upon the principle of gating, which is necessary for the visualisation of correlations in multiparameter data. It is traditionally completed manually, which is time and resource-intensive and subject to possible inconsistency as analysis is subject to individual judgement. Despite these drawbacks, there has been reluctance from some laboratories to move to computational approaches.
Automated gating addresses a lot of the challenges associated with the manual approach (particularly user bias) and offers a range of benefits for researchers, including accelerating data processing, improving reproducibility, and ensuring quality control, speeding up the overall flow cytometry process and ultimately helping pharmaceutical companies to create effective drugs for patients.
Case study: Covid-IP project
In collaboration with Guy’s and St Thomas’ NHS Foundation Trust and the Francis Crick Institute in London, and the European Bioinformatics Institute (EMBL-EBI) in Cambridge, UK, King’s College London (KCL) launched the Covid-IP (Covid–ImmunoPhenotype) project in March 2020, to better understand the immunophenotype of patients infected with SARS-CoV-2.
Immunophenotypes vary greatly across different individuals, giving strong clues as to what mechanisms the human immune system must employ to provide protection from Covid-19, and indicating ways in which it can go wrong, worsening rather than improving the patient’s condition.
The Covid-IP project performed immunophenotyping on blood samples from >120 Covid-19 patients, consisting of eight complementary flow cytometry panels per patient to capture the major populations of immune cells and rare populations, as well as activation markers. This generated thousands of FCS files, requiring significant manpower to organise the analysis, which is manual. Scientists had to create each gate and process each file individually, creating a dual challenge of manpower and the possibility of variability in the data due to manual gating.
As a direct comparison to the manual gating that was carried out on this project, automated pipelines were implemented for flow cytometry analysis (CytoML, Aigenpulse) to automate all steps from data import, QC, gating, statistical analysis and visualisation. This enabled researchers to apply guided algorithms to mimic human gating strategies to the entire dataset without manual intervention.
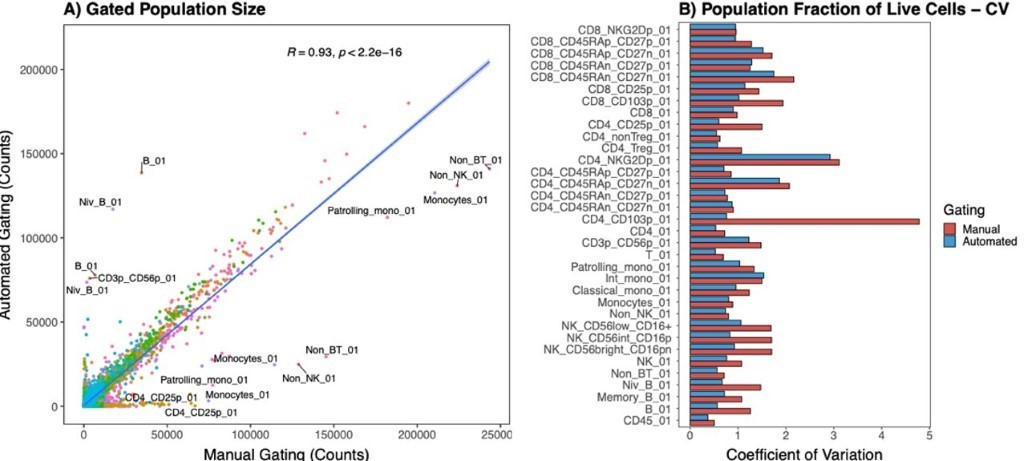
Figure 1: A) Direct comparison of population sizes from 210 samples gated using manual or automated strategies. Colours represent gated populations. Covariance was measured with Pearson’s correlation coefficient, R = 0.93, p-value = 2.2-16. B) Coefficient of variation for gated populations (normalised to live cells), comparing manual and automated gating strategies.
Compared with the KCL manual pipeline, automated processing provided a strong correlation (Pearson’s R = 0.93) (Figure 1a) and reduced variation for each gating step (Figure 1b). The fast processing time reduced the full time equivalent (FTE) from >10 over eight weeks using manual gating, to 1.5 over two weeks using automated gating.
Unlocking insights
The data generated by pharmaceutical R&D holds enormous opportunity for the development of life-changing therapies, but cannot be leveraged without appropriate data analytics to unlock insights and facilitate decision making. For example, more value can be derived from integrating flow cytometry data with both in-house and public single-cell sequencing, proteomics, and transcriptomics data, using platforms that integrate and unify the data, and provide robust machine-learning tools that help uncover novel insights.
Researchers can rapidly explore large data assets to drive development decisions, and use the time saved on laborious data processing for higher value-added tasks.
This is vital for projects such as Covid-IP, where numerous laboratories from different institutions rely on sharing data in real-time and obtaining insights that could aid the diagnosis and treatment of the Covid-19 virus.
About Aigenpulse:
Aigenpulse Limited was founded in May 2016 to develop the unique Aigenpulse IT platform that takes into account the dynamic and evolving nature of research. With offices in Oxfordshire and London in the UK and Boston in the United States, Aigenpulse consists of experts with PhD level qualifications in Life Sciences combined with highly skilled software developers, bioinformatics data scientists, cloud computing engineers and user experience/user interaction designers.
For more information, visit: www.aigenpulse.com.
Topics: Emerging Technologies